New study sheds light on the conservation of the Serine ADP-ribosylation pathway
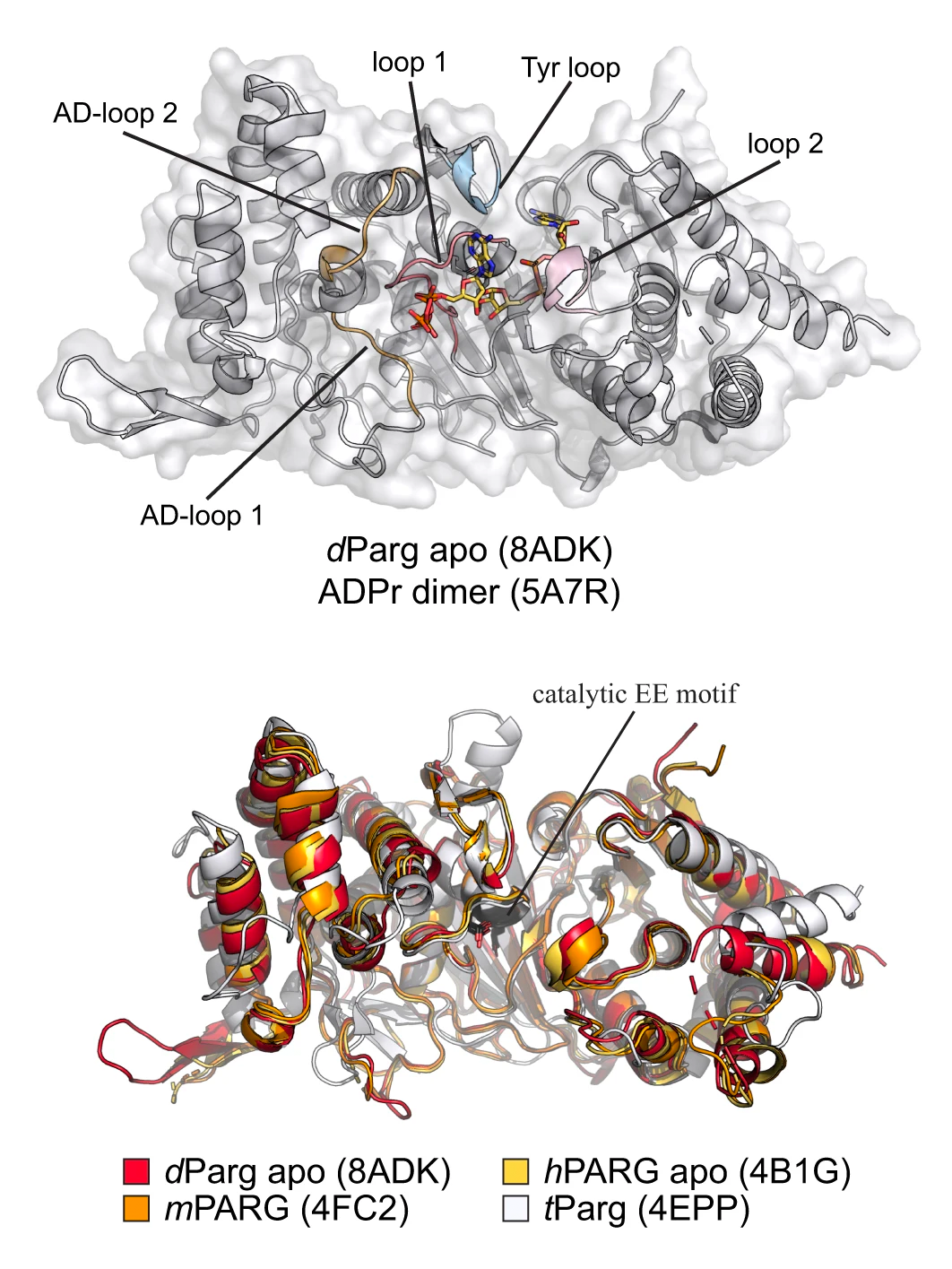
Ribbon representation of dParg, and ribbon representation of structural alignment of PARG domains from different species (Drosophila, red; human, yellow; mouse, orange; T. thermophile, white).
ADP-ribosylation is a post-translational modification involved in a wide variety of cellular processes including the DNA damage response. ADPr chains can be attached by a family of enzymes called PARPs to different amino acids (e.g., glutamate, aspartate, serine, arginine) on target proteins. It was recently discovered that the vast majority of ADP-ribosylation upon DNA damage occurs on serine residues. This activity requires HPF1, an accessory factor which changes PARP1/2 ADP-ribosylating activity towards serine. However, the majority of studies investigating Ser-ADPr has been conducted on mammalian cells, and little was known whether this signalling could be present in organisms outside of Mammalia. The study published in Nature Communications by Fontana et al. addresses this exact question, showing for the first time the conservation of the HPF1 gene across different animal species, and examining the importance of Ser-ADPr in DNA damage response in Drosophila melanogaster.
Previous phylogenic analyses have revealed that fruit flies have conserved the HPF1 gene and are capable of Ser-ADPr. Interestingly, D. melanogaster lack ARH3, a hydrolase which is the only known enzyme capable of removing Ser-ADPr in humans. Despite this, experiments conducted by Fontana et al. on the the ADPr response to DNA damage in Drosophila recapitulated findings from human cells, showing that this pathway is somehow conserved across the two species. Mass spectrometry studies demonstrated a significant overlap between Drosophila ADP-ribosylation targets and human ADP-ribosylation targets, while also identifying 25 novel Drosophila-specific targets. Moreover, it was found that the Drosophila Parg (dParg) enzyme, unlike human PARG, is capable of cleaving the bond between ADPr and its target serine residue, completely removing ADPr from protein. Further studies have shown that this novel activity does not stem from alterations in the catalytic domain of the enzyme. Collectively, the findings of this paper highlight the importance of HPF1 and serine-ADPr pathways in the DNA damage response in Drosophila, and additionally demonstrate the value of this model organism in studying ADP-ribosylation signalling in health and disease.
Osamu Suyari, a postdoc in the Ivan Ahel lab and one of the first authors on the paper, added: “I am really excited to share with you the revelation that Ser-ADP ribosylation is conserved in Drosophila and how it controls the DNA damage response in this species. I thank all the fantastic collaborators. I hope these findings will be able to contribute to the further uncovering of physiological functions of Ser-ADP ribosylation in the future.”
Explore more
Read the paper
Serine ADP-ribosylation in Drosophila provides insights into the evolution of reversible ADP-ribosylation signalling
Ivan Ahel group
The group of Ivan Ahel works on the pathways and protein functions underlying genome stability, and in particular the role of the post-translational protein modification ADP-ribosylation in this process.
Cancer and Genome Stability
Several Dunn School groups study processes that are critical to maintain genome integrity and normal cell growth