Study of Poxviruses Leads to New Treatment Avenues
Poxviruses are large, enveloped, double-stranded DNA viruses that are responsible for a range of diseases in humans and animals. A huge achievement in the fight against poxviruses was the eradication of smallpox, caused by the variola virus. This was declared by the WHO in 1980 after a successful vaccination campaign with vaccinia virus, a related orthopoxvirus. However, despite the continuous efforts, poxviruses still pose a palpable health risk – one that we experienced during the global monkeypox (mpox) epidemic in 2022 caused by monkeypox virus. The drug tecovirimat has been used to combat monkeypox virus during this epidemic, but already more than 20 separate instances of tecovirimat-resistant monkeypox virus have been described, and so additional drugs are needed.
The main aim of the Smith group is studying how the poxviruses evade host defences. Using mass spectrometry, they looked at the proteome of vaccinia-infected cells and discovered a subset of cellular proteins that was being downregulated when compared to uninfected cells. This targeted degradation was hypothesised to be a viral evasion strategy from host proteins that were antiviral, and a detailed study of some of the downregulated proteins showed that in each case they did possess antiviral activity. In a study undertaken at the University of Cambridge, the University of Oxford and the Pirbright Institute, Yiqi Zhao, Yongxu Lu and colleagues took a closer look at one of the identified proteins – TRIM5α – to discover why the virus was trying to get rid of it.
In cells in which TRIM5α was knocked out, vaccinia virus was shown to replicate and spread better, confirming TRIM5α has antipoxvirus activity. It was also discovered that the virus deploys 2 countermeasures to antagonise TRIM5α. Firstly, it induces the proteosome-dependent degradation of TRIM5α by direct binding of a viral protein called C6. Secondly, the virus recruits a cellular protein cyclophilin A (CypA), which antagonises TRIM5α by binding to the same viral capsid protein (called L3) that TRIM5α binds to.
CypA is a target of existing drugs, such as cyclosporine A and several non-immunosuppressive derivatives. Notably, it was shown that these drugs were able to reduce the replication and spread of several different orthopoxviruses, including monkeypox, camelpox, cowpox and vaccinia viruses. This is of particular importance because these drugs target a cellular protein making it difficult for the virus to evolve drug resistance. This is unlike the situation with tecovirimat, which targets a viral protein that can mutate to induce drug resistance.
The interactions between the cellular proteins TRIM5α and CypA, and the viral proteins C6 and L3 that are highlighted in this study, were shown to be conserved across a range of orthopoxpox viruses, including variola virus, suggesting that the CypA-targeting drugs could act against a broad spectrum of poxvirus infections. Despite the potential clinical applications, Prof Smith emphasises that this project started off as a basic science story – one trying to understand the underlying mechanisms of host defences against viruses and viral evasion of these defences. These unforeseen results simply emphasise the importance of investing in basic science research.
Written by Aleksandra Pluta (Murphy lab) @aj_pluta
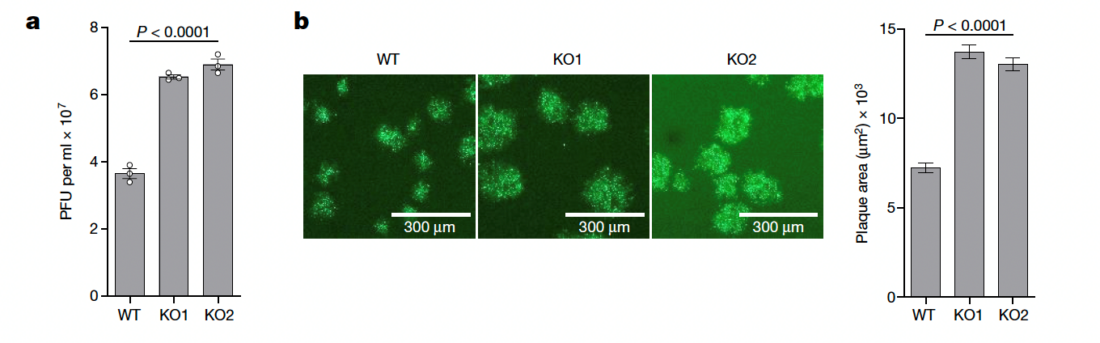
Infectious vaccinia virus titres (a) and plaque images (b) formed on wild-type (WT) and TRIM5α-knockout cells (KO1/KO2) after infection with vaccinia virus.
Read the paper
TRIM5α restricts poxviruses and is antagonized by CypA and viral protein C6
Smith group
Studying the mechanisms by which orthopoxviruses suppress innate immunity
Infection and Immunity
Several Dunn School groups use a range of approaches to investigate antigen presentation and immune regulation during health and disease and study the mechanisms that enable bacterial and viral pathogens to invade and proliferate inside their hosts.